
In 1941, a rose killed a policeman.
Albert Alexander, a 43-year-old policeman in Oxford, England, was pruning his roses one fall day when a thorn scratched him at the corner of his mouth. The slight crevice it opened allowed harmless skin bacteria to slip into his body. At first, the scratch grew pink and tender. Over the course of several weeks, it slowly swelled. The bacteria turned from harmless to vicious, proliferating through his flesh. Alexander eventually had to be admitted to Radcliffe Hospital, the bacteria spreading across his face and into his lungs.
Alexander’s doctors tried treating him with sulfa drugs, the only treatment available at the time. The medicine failed, and as the infection worsened, they had to cut out one of his eyes. The bacteria started to infiltrate his bones. Death seemed inevitable.
But then, on February 12, 1941, Alexander was injected with an experimental drug: a molecule produced by mold.
The molecule was, of course, penicillin. It had been discovered thirteen years earlier but soon abandoned because there didn’t seem to be any way to turn it into an effective drug. In the late 1930s, Howard Florey and his colleagues at the University of Oxford revived the drug and began testing it on mice. They found the penicillin could cure them of infections by killing their bacteria. Florey then gave a dose of penicillin to a woman dying of cancer and found that it wasn’t toxic to her.
Now Florey and his colleagues wanted to see if it could stop an infection in a human being. Alexander, with nothing left between him and death, was their first subject.
“Striking improvement” was how Florey described what happened next. Within a day, Alexander’s infections were subsiding. After a few more days, his fever broke and much of his face cleared up.
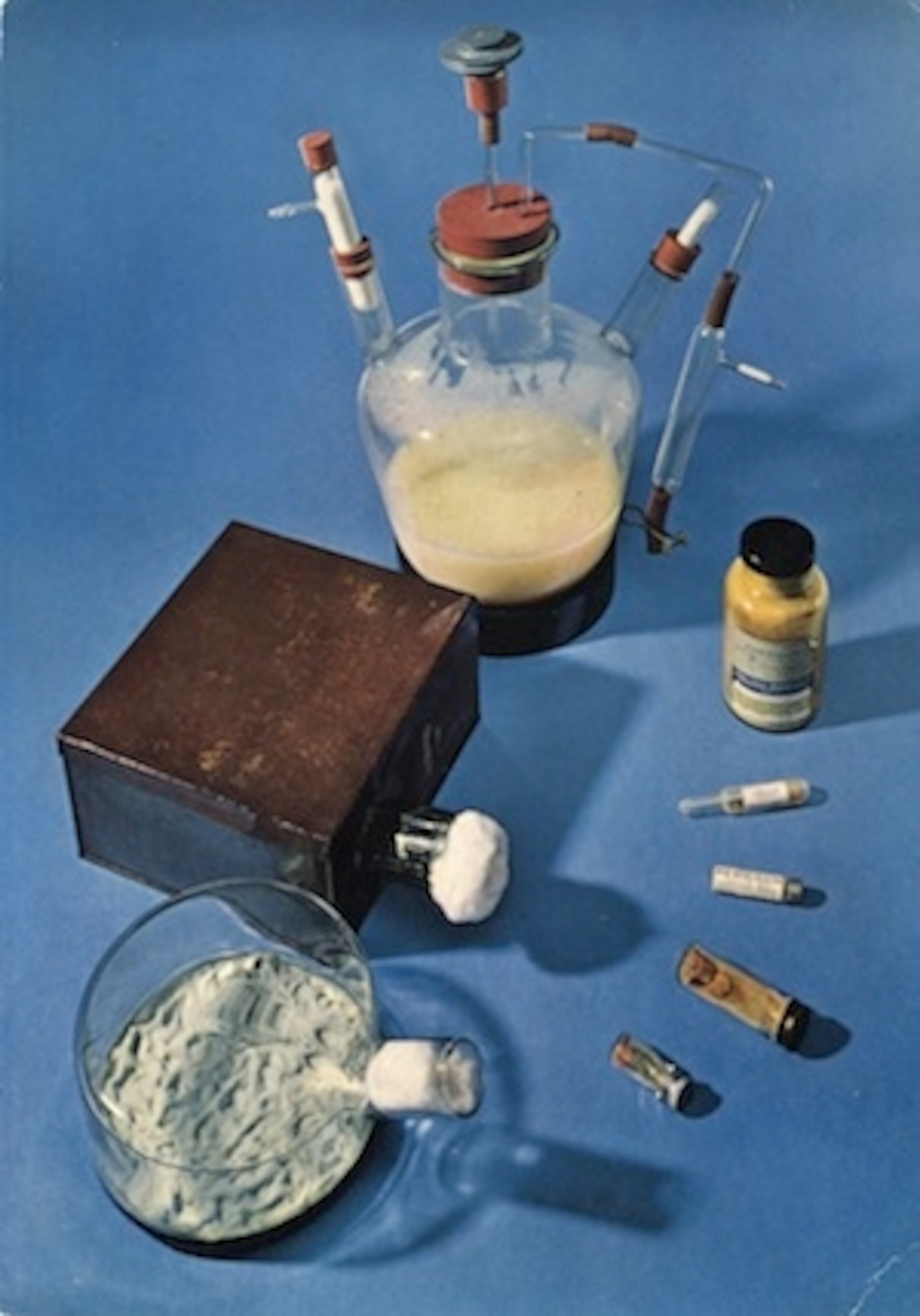

Florey could have saved Alexander’s life, if he hadn’t run out of penicillin after a few days. Nobody but Florey knew how to make the stuff, and his recipe only yielded a tiny amount at a time. To stretch out their supply of penicillin, a member of Florey’s lab would visit the hospital each morning to collect Alexander’s urine. He would carry it back by bicycle to the lab, where the scientists extracted the penicillin that Alexander’s body hadn’t absorbed. Alexander’s doctors then injected the recycled antibotic into Alexander’s arm.
But the salvaging operation didn’t recover enough penicillin to keep the bacteria from growing again. The infection returned and grew worse than before. On March 15, Alexander died. In his final report, Florey called Alexander’s death “a forlorn case.”
It is hard to imagine a time when a scratch could so easily lead to death. Albert Alexander died precisely at the dawn of the Antibiotic Era. Shortly after failing to save Alexander’s life, Florey harvested more penicillin and gave it to another patient at the hospital, a 15-year-old boy who had developed an infection during surgery. They cured him in a few days. Within three years of Alexander’s death, Pfizer was manufacturing penicillin on an industrial scale, packing 7500-gallon tanks with mold, fed on corn steep liquor. In that same year, Selman Waksman, a Rutgers microbiologist, and his colleagues discovered antibiotics made by soil bacteria, such as streptomycin and neomycin.
What made antibiotics so wildly successful was the way they attacked bacteria while sparing us. Penicillin, for example, stops many types of bacteria from building their cell walls. Our own cells are built in a fundamentally different way, and so the drug has no effect. While antibiotics can discriminate between us and them, however, they can’t discriminate between them and them–between the bacteria that are making us sick and then ones we carry when we’re healthy. When we take a pill of vancomycin, it’s like swallowing a grenade. It may kill our enemy, but it kills a lot of bystanders, too.

It’s understandable that few scientists gave this fact much thought in the 1940s, when the lives of people like Albert Alexander hung in the balance. Even if they did wonder about the 100 trillion microbes that live in our healthy bodies–known as the microbiome–they were poorly equipped to investigate them. They could only study bacteria that they could rear in their labs. E. coli thrived outside of the body, sucking in oxygen and feeding on just about any sugar on offer. That’s why scientists now understand E. coli better than any other species on Earth.
But in its natural habitat–the human gut– E. coli is a rare bird. Only about one microbe in a thousand in the gut belongs to the species. The rest of the microbes are far too fussy to survive in just any Petri dish. They need a special balance of gases, acidity, and nutrients. In many cases, they can’t survive unless they’re living alongside other species. Their fussiness has slowed down scientists trying to explore the microbiome. But now that they can fish out the DNA of the microbiome, scientists are beginning to get a sense of the staggering diversity of microbes we harbor.
Each of us is home to several thousand species. (I’m only talking about bacteria, by the way–viruses, fungi, and protozoans stack an even higher level of diversity on top of the bacterial biodiversity.) My own belly button, I’ve been reliably informed, contains at least 53 species. Many of the species I harbor are different than the ones you harbor. But if you look at the kinds of genes carried by those species, our microbiomes look very similar. That’s partly because surviving on a human body requires certain skills, so any species that is going to last long in your lungs, say, will need many of the same genes.
But the similarity speaks to something else. The microbiome keeps us healthy. It breaks down some of our food into digestible molecules, it detoxifies poisons, it serves as a shield on our skin and internal linings to keep out pathogens, and it nurtures our immune systems, instructing them in the proper balance between vigilance and tolerance. It’s a dependence we’ve been evolving for 700 million years, ever since our early animal ancestors evolved bodies that bacteria could colonize. (Even jellyfish and spongeshave microbiomes.) If you think of the human genome as all the genes it takes to run a human body, the 20,000 protein-coding genes found in our own DNA are not enough. We are a superorganism that deploys as many as 20 million genes.
It’s not easy to track what happens to this complex organ of ours when we take antibiotics. Monitoring the microbiome of a single person demands a lot of medical, microbiological, and genomic expertise. And it’s hard to generalize, since each case has its own quirks. What happens to the microbiome depends on the particular kind of bacteria infecting people, the kind of antibiotics people take, the state of their microbiome beforehand, their own health, and even their own genes (well, the human genes, at least). And then there’s the question of how long these effects last. If there’s a change to the microbiome for a few weeks, does that change vanish within a few months? Or are there effects only emerge years later?
Scientists are only now beginning to get answers to those questions. In a paper just published online in the journal Gut, Andres Moya of the University of Valencia and his colleagues took an unprecedented look at a microbiome weathering a storm of antibiotics. The microbiome belonged to a 68-year-old man who had developed an infection in his pacemaker. A two-week course of antbiotics cleared it up nicely. Over the course of his treatment, Moya and his colleagues collected stool samples from the man every few days, and then six weeks afterwards. They identified the species in the stool, as well as the genes that the bacteria switched on and off.
What’s most striking about Moya’s study is how the entire microbiome responded to the antibiotics as if it was under a biochemical mortar attack. The bacteria started producing defenses to keep the deadly molecules from getting inside them. To get rid of the drugs that did get inside them, they produced pumps to blast them back out. Meanwhile, the entire microbiome powered down its metabolism. This is probably a good strategy for enduring antibiotics, which typically attack the molecules that bacteria use to grow. As the bacteria shut down, they had a direct effect on their host: they stopped making vitamins and carrying out other metabolic tasks.
In another intriguing response, the microbes dimmed their immune systems. To defend against invading viruses, bacteria deploy a collection of enzymes that recognize foreign genes and chop them up. As the bacteria dialed these enzymes down, they may have allowed viruses to infect them more easily. In some cases, the invasion led to their death. But in other cases, the viruses may have delivered them useful genes, including genes that let them resist the antibiotics.
Moya and his colleagues found that some types of bacteria were able to survive the onslaught of antibiotics, while others failed. As a result, the overall diversity of bacteria in the man’s gut changed from day to day over the course of his treatment. Before he started taking antibiotics, the scientists identified 41 species in a stool sample. By day 11, they only found 13. Six weeks after the antibiotics, the man was back up to 38 species. But the species he carried six weeks after the antibiotics did not represent that same kind of diversity he had before he took them. A number of major groups of bacteria were still missing.
This long-term disturbance was not unusual. Other scientists have tracked the diversity of the microbiome for many months after people get antibiotics. Even after all that time, the microbiome may not return to its original state. By disturbing our inner ecosystem, antibiotics can affect our own health.
In some cases, for example, antibiotics can make it easier for pathogens to invade. Eric Pamer of Memorial Sloan Kettering Cancer Center and his collegues recently provided a striking demonstration of this effect. They gave mice a single dose of the antibiotic clindamycin. Ninety percent of the diversity in the gut of the mice disappeared and was still gone four weeks after the treatment. The scientists then inoculated the mice with the spores of Clostridium difficile, a particularly nasty pathogen that can cause lethal cases of diarrhea. They invariably got an overwhelming infection, and half of them died within a few days. Pamer could wait as long as ten days after giving the mice antibiotics, and they were still felled by C. difficile. Healthy mice, on the other hand, easily kept the invasion in check.
Antibiotics may also exert subtler, longer-term effects on our health. Matthew Kronman of Seattle Children’s Hospital and his colleagues, for example, recently reviewed the medical records of over a million people. They found that children who took antibiotics were at greater risk of developing inflammatory bowel disease later in life. The more antibiotics they took, the greater the risk. Similar studies have found a potential link to asthma as well.
A study carried out by Dennis Kasper at Harvard hints at how antibiotics can send the immune system off the rails. They reared mice in isolated containers so that they never developed a microbiome. The germ-free rodents developed unusually high levels of an aggressive type of immune cell called an invariant natural killer T cell. If Kasper inoculated baby germ-free mice with a normal microbiome, the T cells remained rare. Antibiotics, the scientists propose, allow the T cells to explode and to run amok.
It’s even possible that long-term antibiotic use may influence how people put on fat. Martin Blaser of New York University and his colleagues carried out an experiment on mice in which they fed the animals antibiotics and then tracked their metabolism. The scientists found that the mice fed with antibiotics developed a higher percentage of body fat than mice that didn’t.*
Antibiotics cause this rise in fat, Blaser and his colleagues argue, by creating long-term changes in the microbiome. The species fostered in the mice produce enzymes that change not just how they break down our food, but also send signals to our own hormones to change the way we store energy from our food.
None of these results would ever lead a doctor to give up on antibiotics altogether. Seventy years after Albert Alexander died, they remain the best tool we have to fight off deadly infections. But we shouldn’t be blasé about them. Doctors often prescribe antibiotics to patients simply on the hunch that they have a bacterial infection. It often turns out that viruses are causing the trouble instead. Many parents are all too familiar with the endless cycle of ear infections and antibiotics. That cycle may take a toll.
There are changes that would help fight against that toll–some that we could make right away, and others that will demand a lot more research before becoming practical. We could become less casual about asking doctors for antibiotics. If DNA-sequencing becomes cheap enough, doctors might become able to diagnose bacteria infections quickly and accurately, so as not to prescribe antibiotics when they can’t help. And when it turns out we are infected, there are other ways to fight bacteria. For a century, some scientists have explored using bacteria-infecting viruses as a weapon against infections, for example.
It might even be possible to fight bacteria with bacteria. Instead of blasting both pathogens and harmless microbes alike, we might tend the microbial garden better and keep down the weeds. The most dramatic example of this gardening is the fecal transplant. Half a million people get C. difficile infections a year, many of which can’t be stopped by antibiotics. Doctors have found that a little stool from a healthy donor can crush these invasions. Fecal transplants may also help against inflammatory bowel disease, by restoring the immune system’s essential partners. Transplants might treat infections elsewhere in the body, from cavities in the mouth to rashes on the skin.
These treatments would do more than tamp down the harmful effects of antibiotics. They’d also help keep antibiotics themselves useful. When Florey first tried out penicillin against Alexander and other patients, he worried that the bacteria might adapt to the drug. It eventually did; for many pathogens, penicillin is now useless because they’ve evolved strong resistance against it. C. difficile and many other pathogens have become resistant against many other antibiotics, too. Developing new antibiotics is essential for stopping this decline, but we will need to use them just as sparingly to slow down evolution’s relentless push.
Otherwise, we may return to a time when roses killed policemen.
[This post emerged from the research I did for a talk I gave last week at Rutgers]
[Images: Grenade, Wikipedia; thorn, macrophile on Flickr via Creative Commons; bacteria, Health Research Board on Flickr via Creative Commons; Penicillin, NIH ]
[Update 12/18 11 am: Fixed Howard Florey’s name.]
[*Upate 12/20 7:20 am: In the original version, I incorrectly stated that the mice’s overall body mass increased. This was only true among female juveniles. By seven weeks, however, there was no increase in body mass–only its composition of fat and lean mass. Thanks to zmil for pointing out my error.]
