
World within a tumour–study shows how complex cancer can be
When I used to work at a cancer charity, I would often hear people asking why there isn’t a cure yet. This frustration is understandable. Despite the billions of dollars and pounds that go into cancer research, and the decades since a war on cancer was declared, the “cure” remains elusive.
There is a good reason for that: cancer is really, really hard.
It is a puzzle of staggering complexity. Every move towards a solution seems to reveal yet another layer of mystery.
For a start, cancer isn’t a single disease, so we can dispense with the idea of a single “cure”. There are over 200 different types, each with their own individual quirks. Even for a single type – say, breast cancer – there can be many different sub-types that demand different treatments. Even within a single subtype, one patient’s tumour can be very different from another’s. They could both have very different sets of mutated genes, which can affect their prognosis and which drugs they should take.
Even in a single patient, a tumour can take on many guises. Cancer, after all, evolves. A tumour’s cells are not bound by the controls that keep the rest of our body in check. They grow and divide without restraint, picking up new genetic changes along the way. Just as animals and plants evolve new strategies to foil predators or produce more offspring, a tumour’s cells can evolve new ways of resisting drugs or growing even faster.
Now, we know that even a single tumour can be a hotbed of diversity. Charles Swanton from Cancer Research UK’s London Research Institute discovered this extra layer of complexity by studying four kidney cancers at an unprecedented level of detail. He showed that the cells from one end of the tumour can have very different genetic mutations to the cells at the other end.
These are not trivial differences. These mutations can indicate a patient’s prognosis, and they can affect which drugs a doctor decides to administer. The bottom line is that a tumour is not a single entity. It’s an entire world.
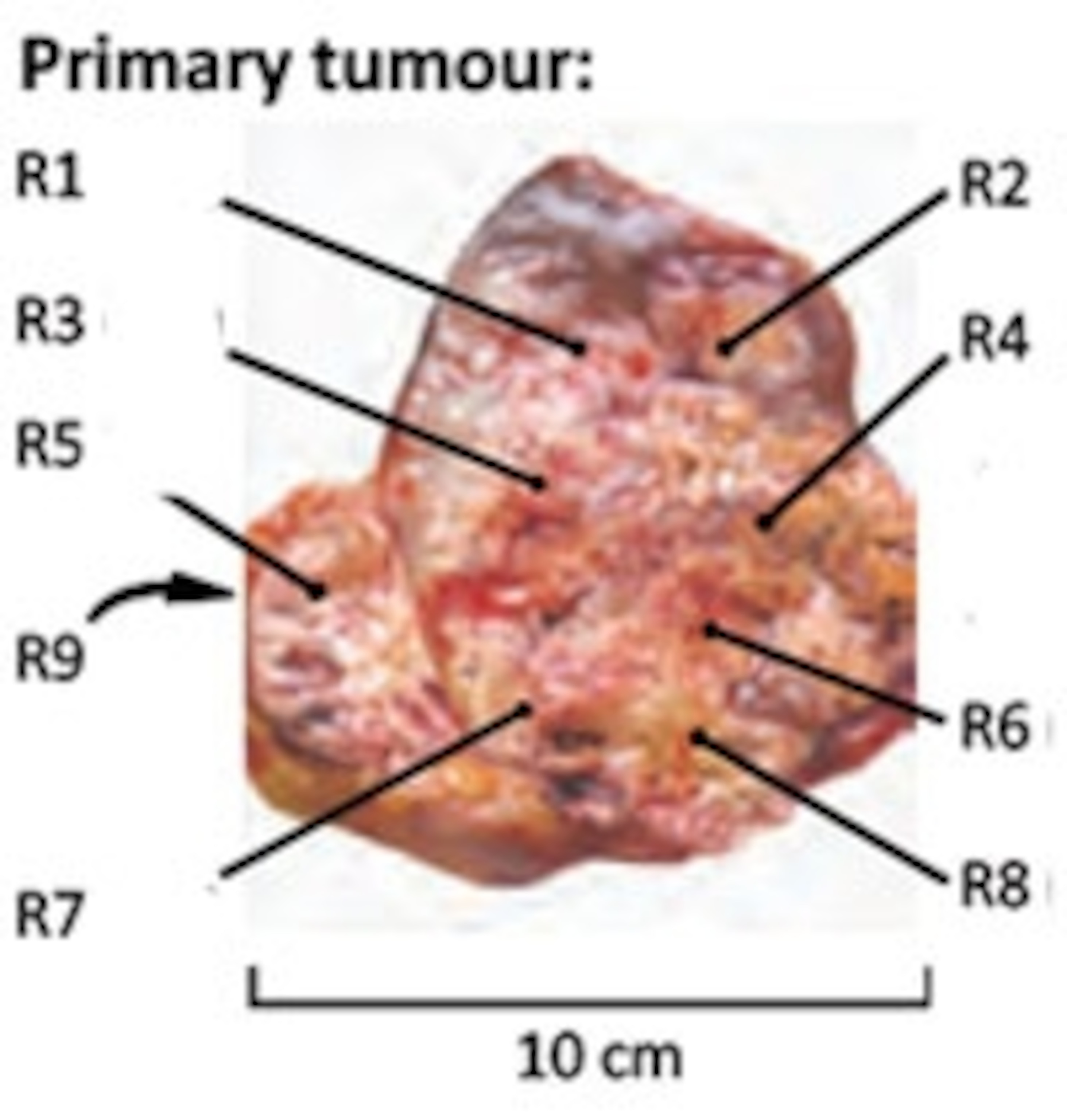
This is one of the tumours that Swanton worked with. It was removed from the kidney of a patient being treated for cancer London’s Royal Marsden hospital. Swanton’s team took nine samples from different parts of the fist-sized mass, as well as other places where it had spread to – the space around the kidney and the patient’s chest.
Swanton found that even the primary tumour was surprisingly varied. He found 128 mutations among the various samples, but only a third of these were common to all of them. A quarter of the mutations were “private” ones – unique to a single sample.
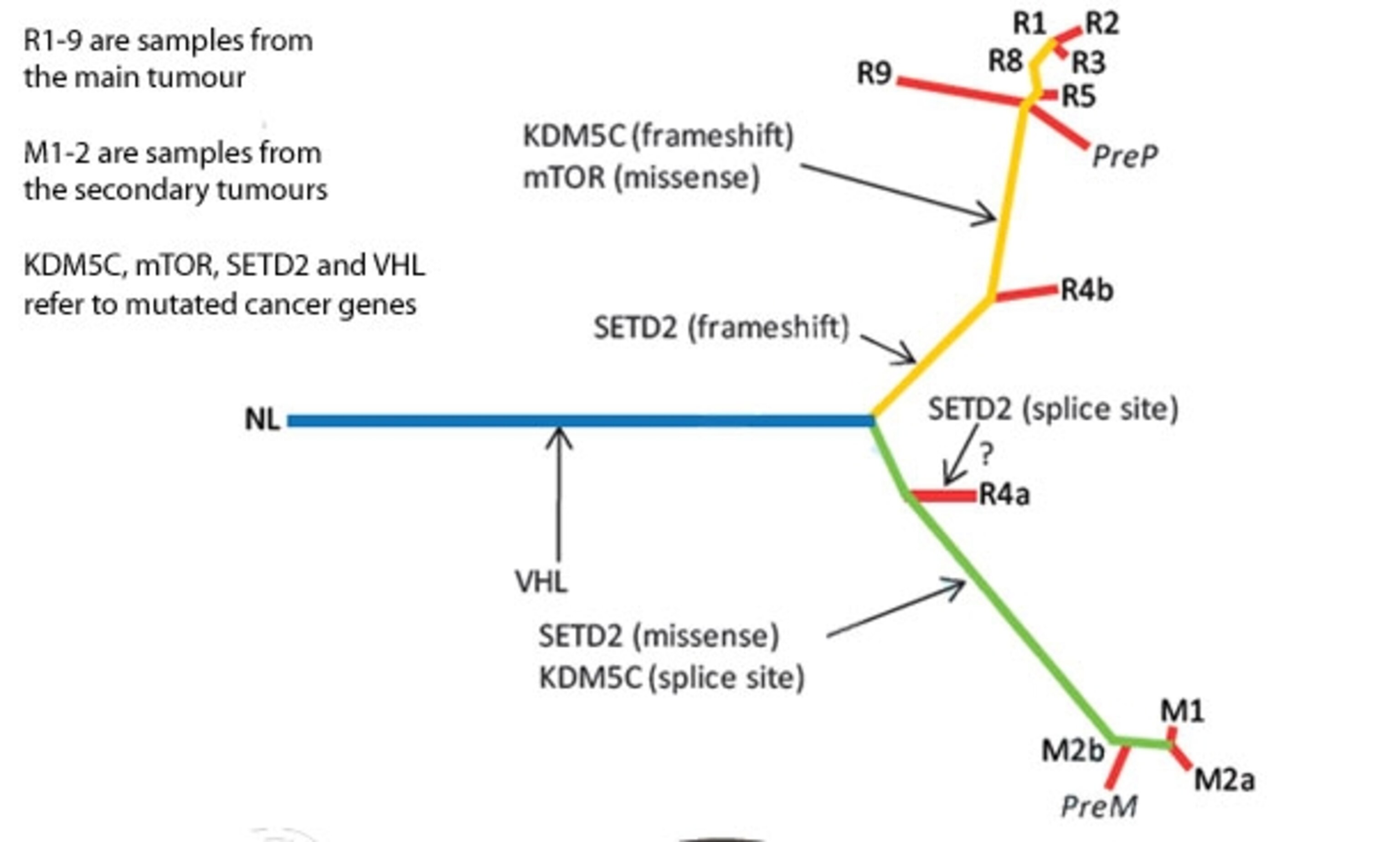
The tumour had also split down two evolutionary lines. One area – part of R4 in the picture – had doubled its usual tally of chromosomes and seeded all the secondary tumours in the patient’s chest. The other branch had spawned the rest of the primary tumour. Even though this tumour looks like a single mass, whose cells all descended from a common ancestor, its different parts arehave all evolved independently of one another.
For example, Swanton found that only one known kidney cancer gene – VHL – was consistently mutated throughout the samples. It was the only one present in the “trunk” of the tumour’s evolutionary tree (the blue line in the image below). The two main branches each picked up their own mutations. However, Swanton also found signs of convergent evolution, where different populations hit on the same adaptations on their own. In this case, three separate parts of the main tumour had disabled the same gene – SETD2 – in different and independent ways.
Three other kidney tumours showed the same degree of bewildering variety and convoluted history.
Darryl Shibata, a cancer geneticist from the University of Southern California, compares Swanton’s approach to the standard medical practice of taking a patient’s history. “By listening to a patient, a wise MD can just about predict what they will find on lab tests,” he explains. “An MD who simply orders tests will get back a baffling list of abnormal findings.” Likewise, reconstructing the evolutionary history of a tumour can make sense of the baffling array of mutations it contains. “Nothing in biology makes sense except in the light of evolution,” adds Shibata, quoting the famous biologist Theodosius Dobzhansky.
Swanton’s study has big implications for the way we study cancers, and could help to explain many of the failures and difficulties that clinicians have faced.
Consider the biomarker problem. For decades, scientists have tried to find molecules that could give away the presence of a cancer, how aggressive it is, what its weaknesses are, and more. They’ve identified tens of thousands of candidates, but only a hundred or so are actually used in practice.
Swanton thinks that this is because people often look for biomarkers after taking a single biopsy of a tumour. That’s potentially misleading. The biomarker might only be relevant to a tiny bit of a tumour, rather than the whole thing. For example, in one of his patients, Swanton found genetic signatures associated with both good and poor prognosis in different parts of a single tumour. “To identify a robust biomarker, that biomarker must be present in all biopsies of the same tumour,” he explains. “A biomarker in the trunk is going to be better than a biomarker in the branches.”
The same goes for treatments. “Personalised medicine” is the next big thing in cancer. This term refers to drugs and treatments that target the specific milieu of faults within an individual patient’s tumour. But of course, a tumour isn’t a single thing. If you target mutations on the branches, you only hit parts of the tumour. Even worse, you leave a big “evolutionary reservoir” of unaffected cells, which can adapt to the drugs you’re using and grow anew. A tumour is like the mythical hydra. If you cut off a head, more will grow back; you need to kill the entire body.
This could explain why many current treatments eventually stop working. “We deal with patients, day in and day out, whose disease initially benefits from chemotherapy but who develop resistance over time,” says Swanton. “The only way you can fathom how a tumour can be one step ahead of the clinician at all times is through this type of diversity and Darwinian selection.” To get around this problem, scientists will need to find the “driver mutations” that are present in all parts of a tumour, in its trunk rather than its branches. These will present the most inviting targets for drugs.
Swanton’s results could also explain why kidney cancer patients fare better if surgeons remove their main tumour, even if their cancer has spread. “It may be that by removing the evolutionary sink of diversity, you improve a patient’s outcomes,” says Swanton. “There’s less material for the tumour to adapt with in response to environmental pressures.”
The study could also cause problems for scientists who are trying to decipher “cancer genomes”. By identifying the full suite of mutations behind any individual tumour, they hope to better understand how cancers develop and what their weaknesses are. But these studies have taken a single biopsy from each patient. And as we now know, that’s often not enough.
The problem is that taking many samples from a tumour is very difficult, and sequencing those extra samples would be very expensive. The alternative is to sequence the one sample over and over. It’s not ideal, but it would reveal small populations of cells that might be distinct from their neighbours and would otherwise stay hidden. “This is likely to be the workhorse way we get at the problem,” says Andy Futreal from the Wellcome Trust Sanger Institute, who was involved in Swanton’s study.
Even this approach has drawbacks. More sequencing means more expense, and cancer genome projects are already costly affairs. “We may need to sacrifice overall numbers to really understand how deep the rabbit hole is,” Futreal says.
If this is all starting to sound very depressing, there is a glimmer of optimism. Even though cancer is maddeningly complicated, survival rates are still going up. At the moment, half of all cancer patients will survive for at least five years after their diagnosis. Cancer seems like an unbeatable adversary, but we are beating it. Studies like Swanton’s show that victory will not come in one decisive strike, but through a thousand shallow cuts. It will take a lot of time and effort.
Consider what Swanton needed to do to complete this study. He needed to piggyback on a clinical trial of kidney cancer patients and work very closely with the surgeons to ensure that all the samples were collected in the right way. It took four years just to get enough samples, and the man who resected all the tumours, Tim Christmas, passed away during that time. Swanton says, “He was an amazing guy and well recognised for being one of the most talented surgeons in Europe. We see this work as a legacy for him.”
Once he had the samples, Swanton threw technique under the sun at them. He sequenced their genes to look for individual mutations – the equivalent of typos in a book. He checked the activity of different genes. He looked for big changes in the structure or number of chromosomes. Most similar projects typically focus on the mutations, but that’s like looking for typos when pages of a book have been smudged, or entire chapters have been rearranged.
“We used every possible genomics technique available,” Swanton says. “Even then we are only scratching the surface of the complexity within each cancer. It has been said you could have one whole scientific institute working on deciphering the genomic events in one tumour.” Even dealing with the data from the study was a challenge. “Our servers were clogged up for two to three months,” Swanton says. “We’re going to have to make a big investment on computing and bioinformatics expertise.”
That’s the type of project we need to really understand cancer’s true nature. Swanton’s study is just the beginning. So far, he has analysed four tumours. “We need to repeat our study in tens or hundreds of tumours, not just four,” he says.
Citation: NEJM, tbc.
Image by Katerha