
Shakespeare’s Sonnets and MLK’s Speech Stored in DNA Speck
When Nick Goldman first opened the package, he couldn’t quite believe that it contained anything at all, much less all of Shakespeare’s sonnets. The parcel had come from a facility in the US and arrived at the European Bioinformatics Institute in the UK, in March 2012. It contained a series of small plastic vials, at the bottom of which were… apparently nothing. It was Goldman’s colleague Ewan Birney who showed him the tiny dust-like specks that he had missed.
These specks were DNA, and they contained:
- All of the Bard’s 154 sonnets.
- A 26-second clip of Martin Luther King’s legendary “I have a dream” speech
- A PDF of James Watson and Francis Crick’s classic paper where they detailed the structure of DNA
- A JPEG photo of Goldman and Birney’s institute
- A code that converted all of that into DNA in the first place
The team sent the vials off to a facility in Germany, where colleages dissolved the DNA in water, sequenced it, and reconstructed all the files with 100 percent accuracy. It vindicated the team’s efforts to encode digital information into DNA using a new technique—one that could be easily scaled up to global levels. And it showed the potential of the famous double-helix as a way of storing our growing morass of data.

A better format
DNA has several big advantages over traditional storage media like CDs, tapes or hard disks. For a start, it takes up far less space. Goldman’s files came to 757 kilobytes and he could barely see them. For a more dramatic comparison, CERN, Europe’s big particle physics laboratory, currently stores around 90 petabytes of data (a petabyte is a million gigabytes) on around 100 tape drives. Goldman’s method could fit that into 41 grams of DNA. That’s a cupful.
DNA is also incredibly durable. As long as it is kept in cold, dry and dark conditions, it can last for tens of thousands of years with minimal care. “The experiment was done 60,000 years ago when a mammoth died and lay there in the ice,” says Goldman. Readable DNA fragments have been recovered from such mammoths, as well as a slew of other prehistoric creatures. “And those weren’t even carefully prepared samples. If you did that under controlled circumstances, you should be good for more than 60,000 years.”
(For those of you wondering if the information would mutate, it can’t. It’s not inside a living thing, and not being copied. It’s just the isolated non-living molecule.)
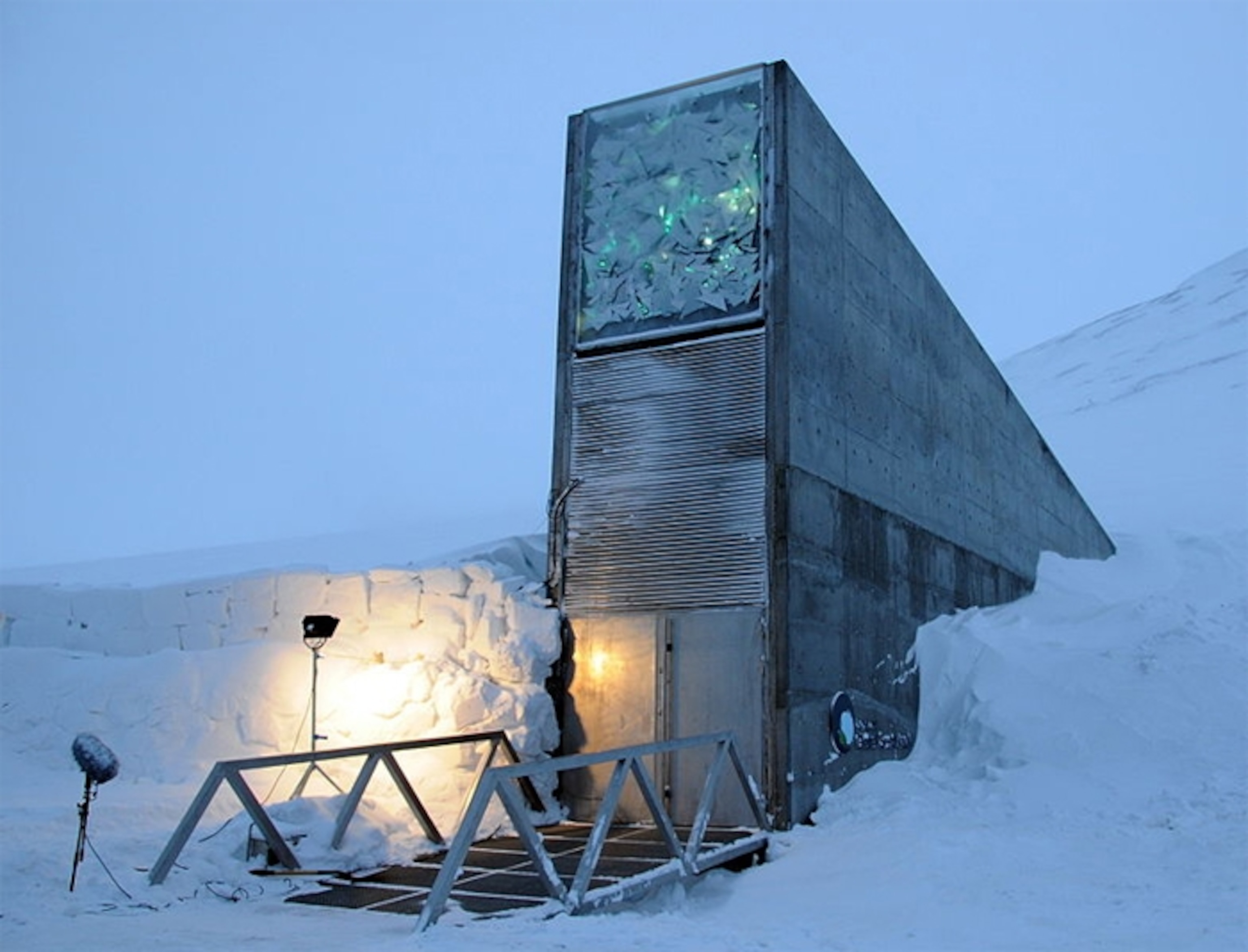
And using DNA would finally divorce the thing that stores information from the things that read it. Time and again, our storage formats become obsolete because we stop making the machines that read them—think about video tapes, cassettes, or floppy disks. That’s a faff—it means that archivists have to constantly replace all their equipment, and laboriously rewrite their documents in the new format du jour, all at great expense. But we will always want to read DNA. It’s the molecule of life. Biologists will always study it. The sequencers may change, but as Goldman says, “You can stick it in a cave in Norway, leave it there in a thousand years, and we’ll still be able to read that.”
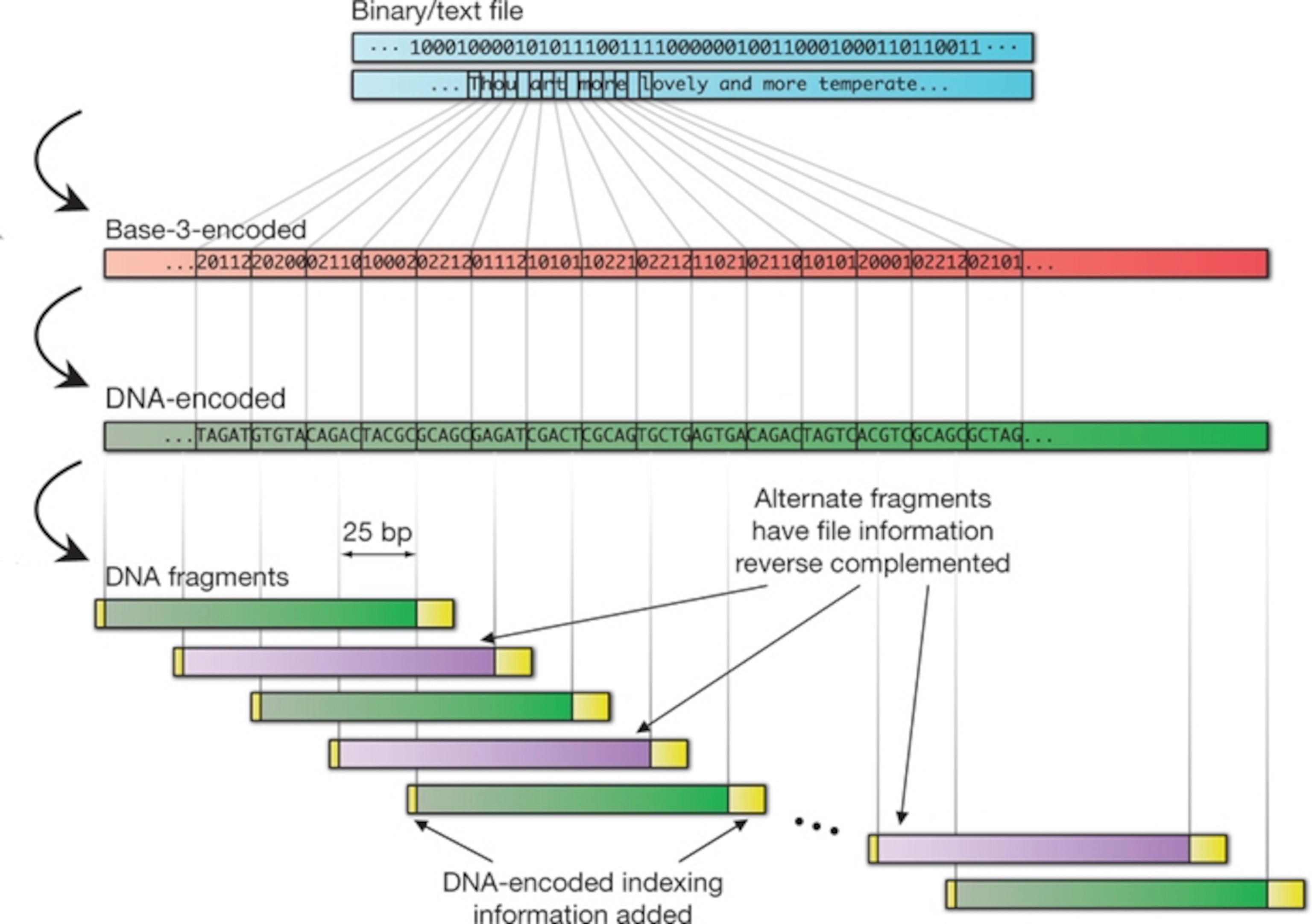
The code
DNA has a proven track record for storing information. It already stores all the instructions necessary to build one of you, or a giraffe, or an oak tree, or a beetle (oh so many beetles). To exploit it, all you need to do is to convert the binary 1s and 0s that we currently use into the As, Gs, Cs and Ts of DNA.
A Harvard scientist called George Church did exactly that last year. He used a simple cipher, where A and C represented 0, and G and T represented 1. In this way, he encoded his new book, some image files, and a Javascript programme, amounting to 5.2 million bits of information
Goldman and Birney have encoded the same amount, but with a more complex scheme. In their system, every byte—a string of 8 ones or zeroes—is converted into five DNA letters. These strings are designed so that there are never any adjacent repeats. This makes it easier for sequencing machines to read and explains why they had a far lower error rate (that is, none) compared to Church’s method.
Using their cipher, they converted every stream of data into a set of DNA strings. Each one is exactly 117 letters long and contains indexing information to show where it belongs in the overall code. The strings also overlap, so that every bit is covered by four separate strings. Again, this reduces error. Any mistake would have to happen on four separate strings, which is very unlikely.
Accuracy aside, Goldman’s coding system has a more fanciful advantage—it should be apocalypse-proof. Let’s get a bit fanciful: Imagine that there’s a calamity that wrecks human civilisation, creating a huge discontinuity in our technology. The survivors rebuild and eventually relearn what DNA is and how to decode it. Maybe they find some of these stores, locked away in a vault. “They’d quickly notice that this isn’t DNA like anything they’ve seen,” says Goldman. “There are no repeats. Everything’s the same length. It’s obviously not something from a bacterium or a human. Maybe it’s worth investigating. Of course you’d need to send some sort of Rosetta stone to tell people how to decode the message…”

Scaling up
Goldman calculated that this method could be feasibly scaled up to cover all of the world’s data (which currently stands at around 3 zettabytes—3 million million gigabytes). For now, the big problems are cost and speed. It’s still expensive to read DNA, and really expensive to write it. The team estimate that you would pay $12,400 to encode every megabyte of data, and $220 to read it back, based on current costs. But those costs are falling exponentially, far faster than those of other electronics.
If you use DNA, you face a steep one-time cost of writing the data. If you use other technologies, you face the recurring costs of having to re-write the data into whatever new format has arrived. It’s the ratio between these two prices that drives the economics of DNA storage.
At the moment, DNA only becomes cost-effective if you want to store things for 600 to 5000 years—that’s the threshold where the one-time cost outweighs all the constant re-writing. But if the price of writing DNA falls by 100 times in the decade, as it assuredly will, then DNA becomes a cost-effective option for storing anything beyond 50 years. “Maybe you’d store your wedding videos,” says Goldman.
DNA technology is also getting faster, but for now, it only makes sense to use it for data that you want to keep for a very long time but aren’t going to access very often.
CERN’s a good example. By 2015, the Large Hadron Collider will be collecting around 50 to 60 petabytes every year—that’s a lot of tape! They also have to migrate their entire archives to new media every four to five years, to save space and avoid the cost of maintaining old equipment. And although people rarely use old data, it has to be kept for at least 20 years, and probably even longer. DNA could be a perfect means of storing these archives (although CERN’s senior computer scientist German Cancio tells me that it will still have to be read and verified every 2 years).
Reference: Goldman, Bertone, Chen, Dessimoz, LeProust, Sipos & Birney. 2013. Towards practical, high-capacity, low-maintenance information storage in synthesized DNA. Nature http://dx.doi.org/10.1038/nature11875