
Holy SARS Origins, Batman!
It’s been more than ten years since the first cases of SARS (severe acute respiratory syndrome) were identified in southern China. It spread to four continents and infected at least 8,200 people before heroic efforts finally brought its progress to a halt. Since then, scientists have sequenced the genome of the virus behind the disease—a coronoavirus called SARS-CoV—and teased apart its infectious tricks. But one lingering question remained: Where did it come from?
At first, the answer seemed straightforward. When investigators swept Chinese animal markets, they found SARS-CoV in palm civets. The animals were quickly culled in their thousands, but later studies showed that their wild or farmed relatives don’t actually carry the virus. They may have acted as a stepping stone for SARS-CoV, but they weren’t its natural reservoir.
Then in 2005, two teams of scientists found that Chinese horseshoe bats harbour a wide range of SARS-like coronaviruses, and SARS-CoV itself belongs to f this diverse family. It seemed that bats, not civets, were the source of SARS—a conclusion that was supported by several later studies.
“But there were doubters,” says Peter Daszak from EcoHealth Alliance, who was involved in one of these studies. “We were among them.”
There were a few niggling problems. The genes of the new coronaviruses had been sequenced but no one had managed to grow one in a laboratory and take a look at it. None of them looked like they could attack the same target that SARS-CoV does—a human protein called ACE2. And although they were all similar to SARS-CoV, none was a good enough match to actually be a direct ancestor. “It would have taken around 100 mutations to turn those into something that could directly infect people,” says Daszak. “That’s not something you’d expect from a bat in a wildlife market.”
The team have now quelled these doubts. They’ve been spending the last few years travelling to 20 countries and searching for new viruses among groups of mammals that are notorious as virus-carriers. Bats are very much on the list, and Chinese bats in particular.
Led by Xing-Yi Ge, Jia-Lu Li and Xing-Lou Yang, the team spent a year in the southern city of Kunming, repeatedly taking faecal samples and anal swabs from a colony of horseshoe bats. They identified seven strains of coronaviruses, including two new ones that were 95 percent identical to SARS-CoV. That’s a far closer match than any virus thus far discovered.
The team even manage to isolate one of the new viruses and grow it in the lab. It could infect human lung cells, as well as cells from pigs and horseshoe bats. And it used the ACE2 protein as a gateway for breaking into its hosts. It’s the most SARS-like virus ever found.
“That’s remarkable,” says Thomas Ksiazek, a virologist who was the lead author on the paper that first described SARS-CoV. “They’re the first ones to make an isolate, which authenticates the whole thing.”
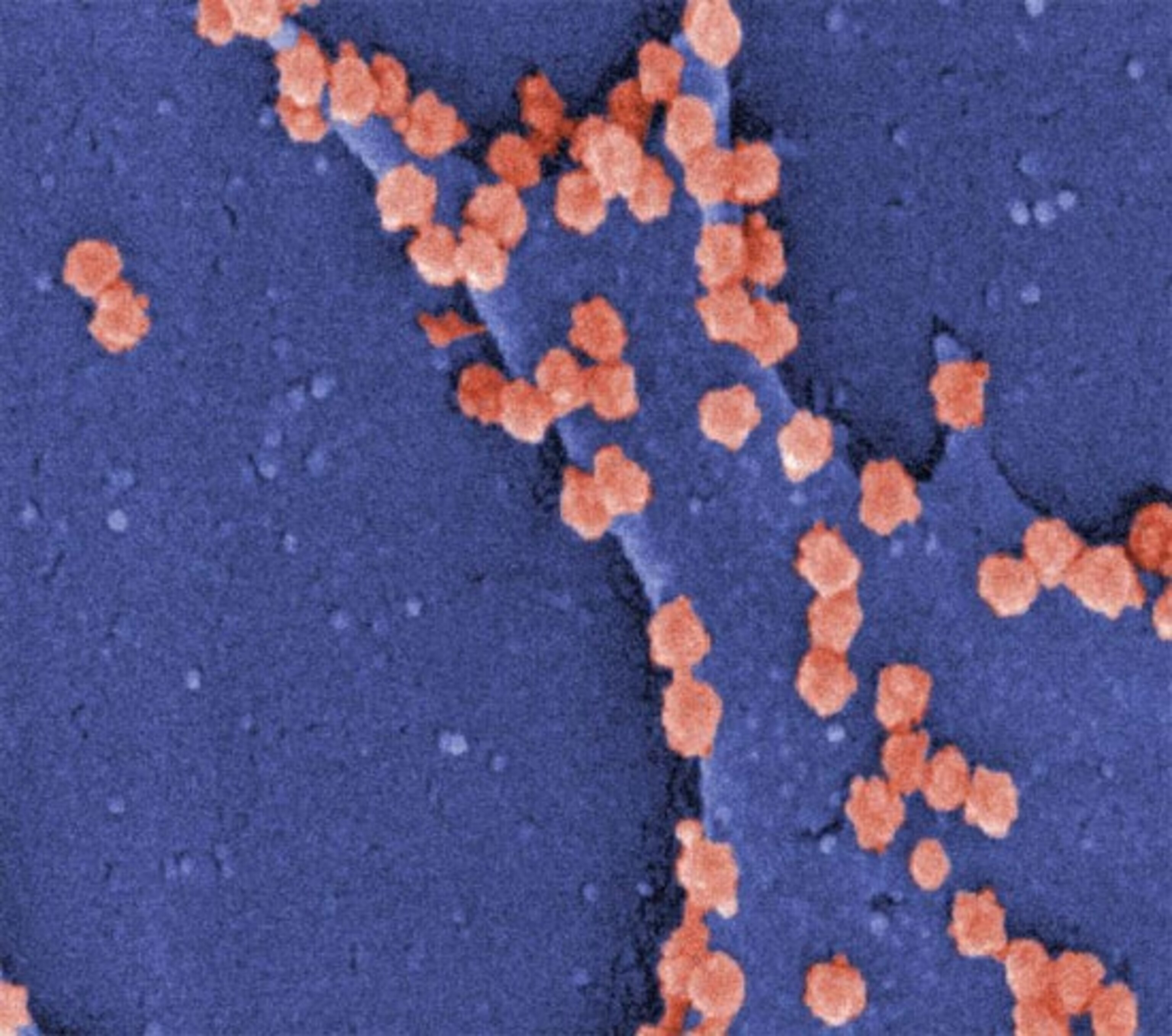
This is the strongest evidence yet that SARS-CoV originated in bats. “Maybe a bat with this virus or something similar got into a market, along with dozens of mammals it had never got into contact with before. Civets, ferret-badgers, rabbits, you name it,” says Daszak. Those other species could have acted as stepping stones on the virus’s path from bats to humans, but since the new strains can stick to ACE2 and infect humans cells, they may have been able to jump directly. “This tells me that we didn’t need civets, or even markets, for SARS to emerge,” says Daszak.
Ksiazek thinks that’s unlikely. “If you look at bat-borne diseases, instances of direct transmission to humans aren’t common, with exceptions like Ebola and Marburg,” he says. Likewise, Christian Drosten from the University Hospital at Bonn still believes that civets were an intermediate host. “That doesn’t make the paper less valuable,” he says. “It means that not every virus sitting in bats is dangerous, but there are some which probably are.
Coronaviruses were mostly neglected until SARS came along, but they seem to be an increasing problem. There’s SARS-CoV itself. There’s the new virus that’s responsible the emerging threat of Middle East Respiratory Syndrome (MERS). And there are goodness knows how many undiscovered strains, which could infect humans. “They’re very evolvable,” says Daszak. “They don’t correct errors in their genome when they make copies of themselves, so you get a lot of mutations, which allows them to lock onto new receptors in new hosts. They’re very good at jumping hosts.”
The team are now checking people who work in Chinese wildlife markets and farms to see if they’ve already been infected by the new SARS-like viruses. And they’re starting pragmatic programmes to minimise the risk of future spillovers. No matter your feelings on China’s wildlife trade, it’s a cultural phenomenon that’s not going away any time soon. So rather than calling for bans, the team are trying to educate farmers about the species most likely to harbour deadly viruses, and steer them towards safe choices. “We know a guy who farms bamboo rats, while other farmers bring bats in from the wild,” says Daszak. “We’d suggest that bamboo rats are a better alternative to bats right now.”
They’re also pushing for bigger projects to identify undiscovered viruses. “It’s pretty sad that it’s taken so long to find these [new corona]viruses,” says Daszak. “We as a species are pretty pathetic about getting ready for pandemics. We sit here and wait for them to emerge, and then we say, Wow, where did that come from?”
Sick of being caught on the backfoot, Daszak and his colleagues are spearheading a new approach to identify every single mammalian virus before they have a chance to spread to humans. They reckon that there are around 320,000 of them, and that it would take USD $1.4 billion to find them all. That’s a trivial cost compared to the price of a pandemic—SARS alone is estimated to have cost USD 16 billion.
To continue reading about this new approach, see my piece for The Scientist. And for more about the SARS story, do read David Quammen’s wonderful book, Spillover.
Reference: Ge, Li, Yang, Chmura, Zhu, Epstein, Mazet, Hu, Zhang, Peng, Zhang, Luo, Tan, Wang, Zhu, Cramer, Zhang, Wang, Daszak & Shi. 2013. Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor. Nature http://dx.doi.org/10.1038/nature12711