Some of my friends are sporting wristbands these days that keep track of their bodies. Little computers nestled in these device inside record the steps they take each day, the beats of their heart, the length of their slumbers. At the end of each day, they can sit down at a computer and look at their data arrayed across a screen like a seismogram of flesh.
I got one of these devices as a gift recently. But as much as I enjoy wasting time with technology, I just didn’t care enough to put it on my wrist. I already know that I should run more, walk more, stand more, and avoid sitting in front of monitors more. I don’t need granular data to remind me of that.
But as I read the journal Genome Biologythe journal Genome Biology today, I decided that someday I might surrender to the Quantified Self movement. I’ll just have to wait till I can track my trillions of microbes from one day to the next.
Thanks to the falling cost of sequencing DNA, it’s now possible for us to survey the thousands of species that live in our bodies. A couple years ago, for example, I found out that I have 58 species in my bellybutton. But all I knew was that there were 58 species in my bellybutton at one point in time–that moment I swiped a Q-tip around my navel. But everything we know about bacteria tells us that our inner ecosystems can change swiftly. My bellybutton may be remarkably different today than it was when I put a Q-tip in it.
Eric Alm, a biologist at MIT, and a graduate student of his named Lawrence David decided to plumb this change by tracking a year in the life of their microbiomes. Each day, they saved some of their stool, and later, they extracted DNA from it to figure out which species of bacteria were living in their guts. David also spat some of his saliva into a tube each day so that he could compare how his microbiome changed in his gut compared to his mouth.
Even though their study only involved two people, it was still very much a Big Data project. And one of the major challenges of any Big Data project is to visualize the results in a useful way. A number on a wrist watch won’t cut it. Between Alm and David, they and their colleagues identified thousands of species of microbes. Most were rare, while a few hundred made up the majority of bugs in their bodies. Some species showed up briefly and vanished; others lingered all year.
Here’s one way to look at their microbiomes. It shows David’s saliva. Each band represents one of the dominant species (or operational taxonomic units). Species belonging to the same lineage (a phylum) have different shades of the same color.
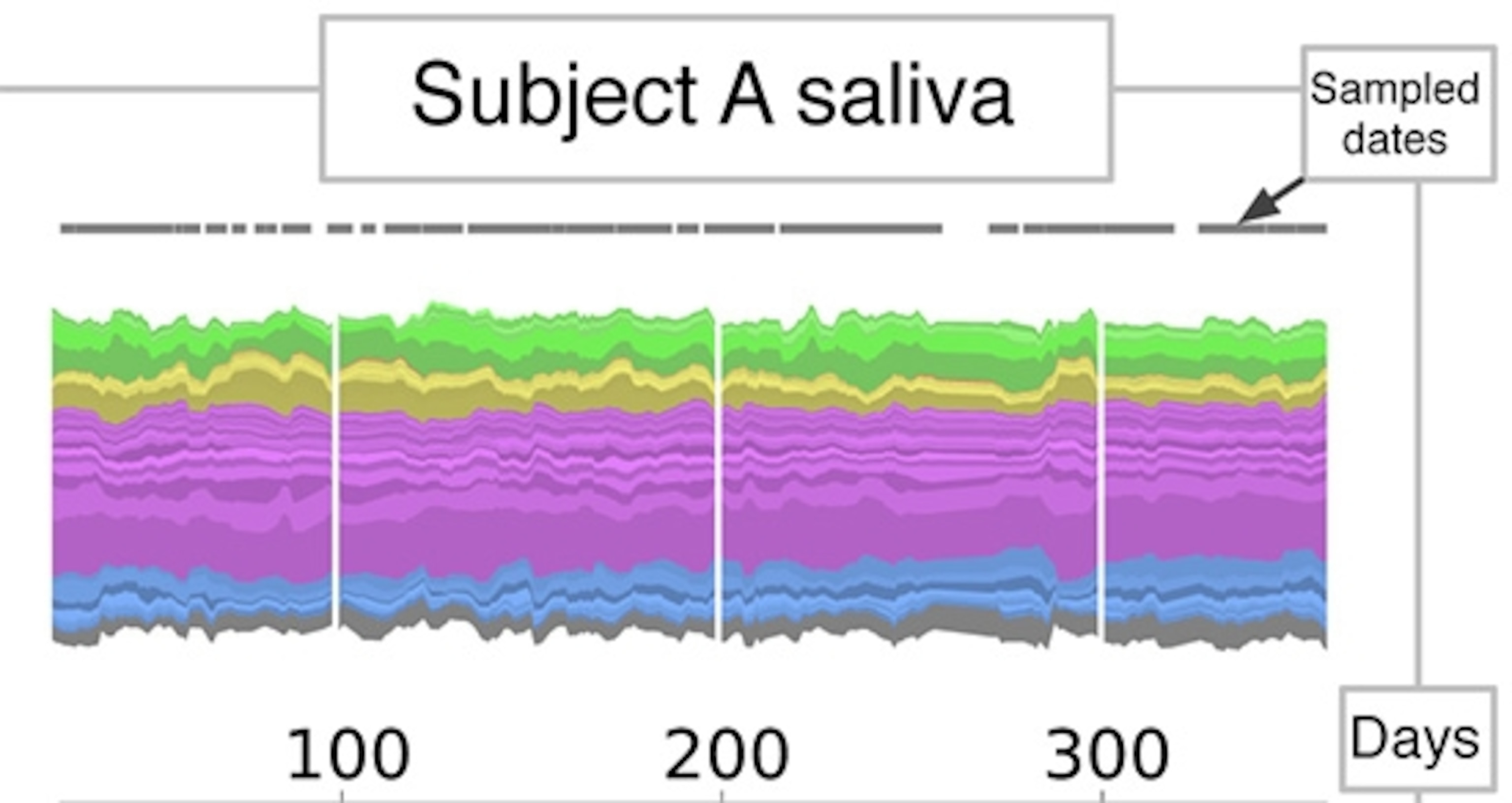
It’s pretty stable over time, but shows some interesting flickers.
Now here’s David’s gut. A different set of species, and more turbulent from time to time, especially around day 100.
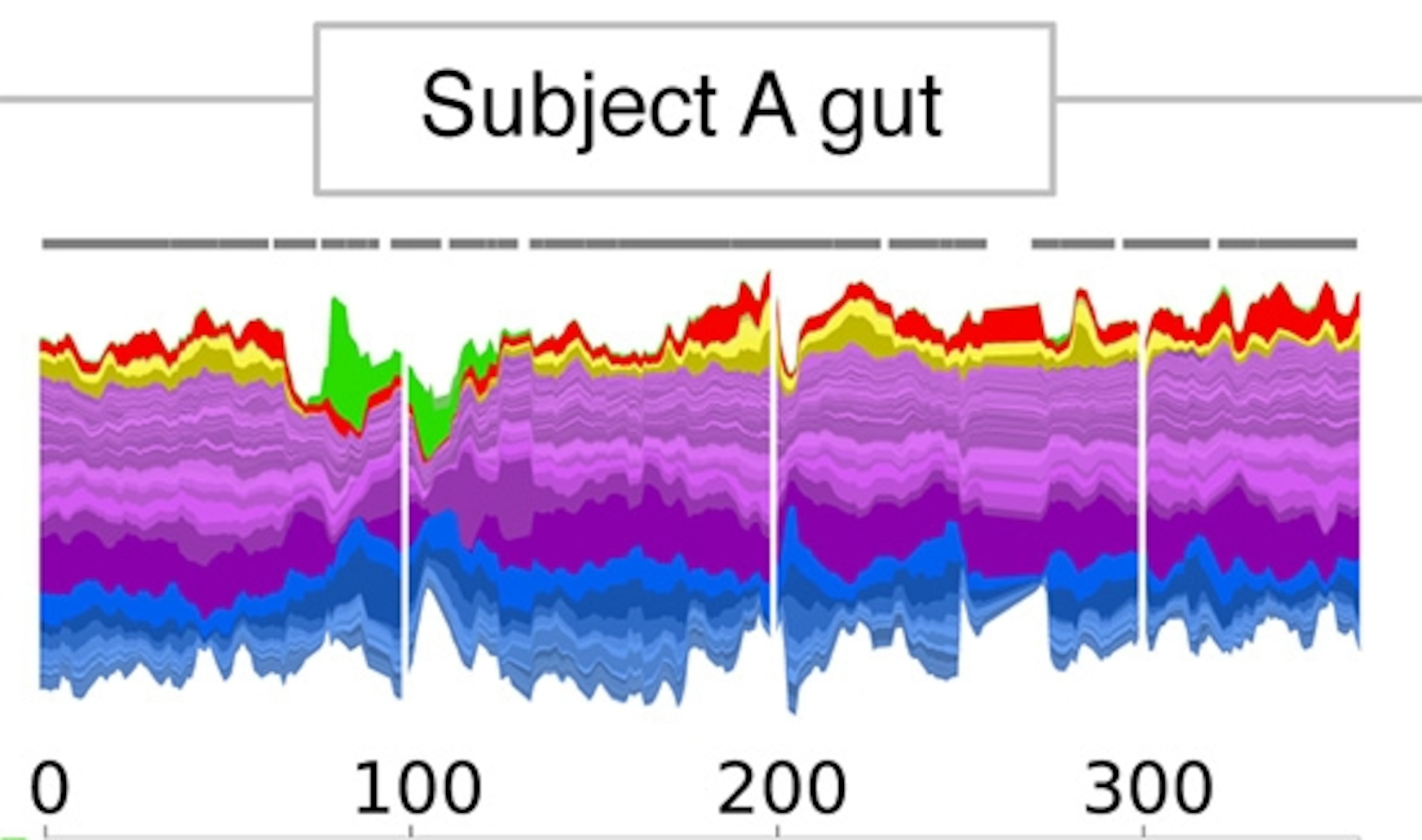
Something’s going on. One way to see what’s happening is to look at the microbiome in a different way. The lower figure shows the changes in each species, with drops in blue and rises in red. As you can see, some species became scarce about a third of the way into the experiment and then returned later. Likewise, other species flourished at the same time and then faded away.
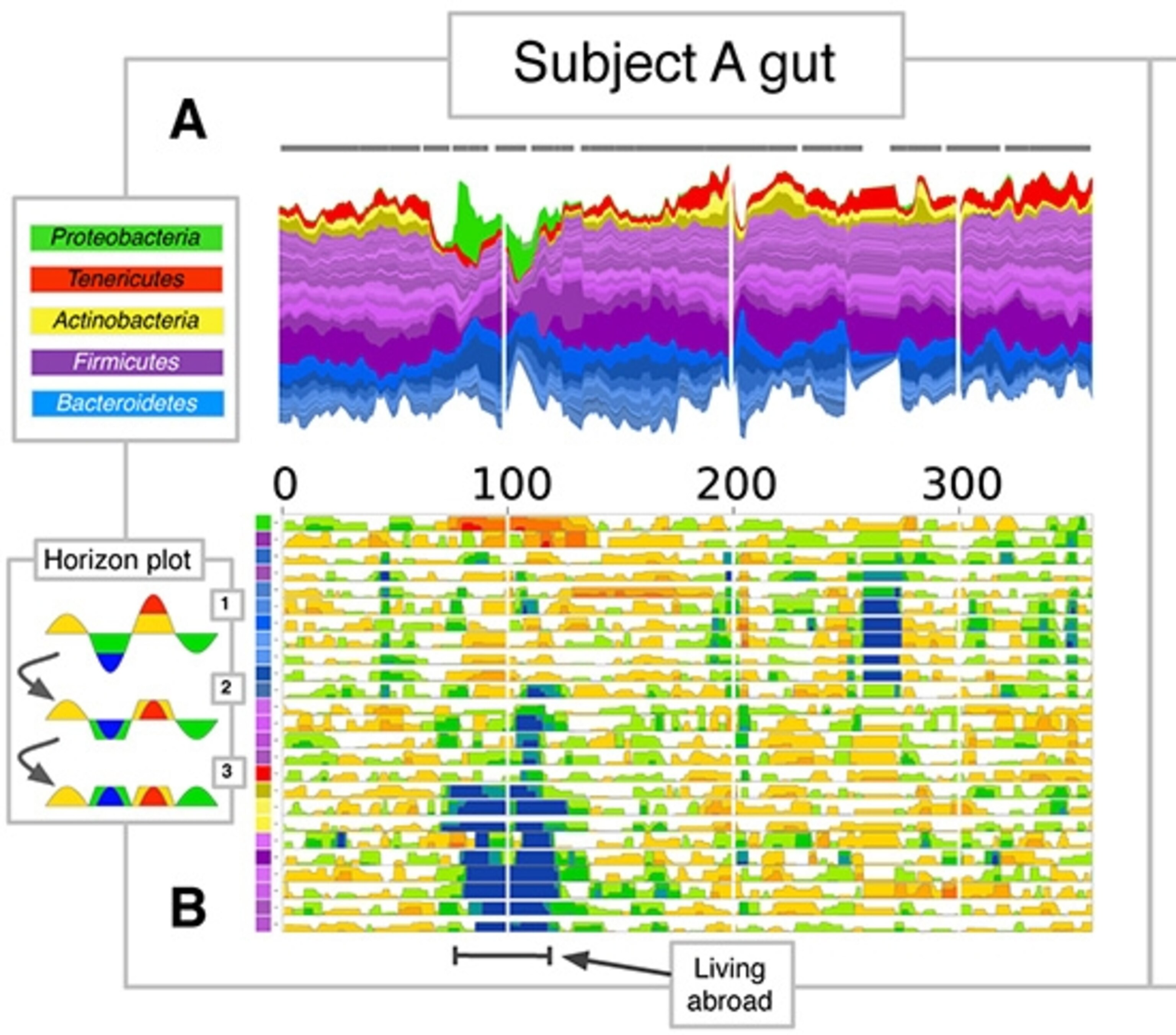
It just so happens that this was when David went to Bangkok for a few weeks, where he had a couple bouts of diarrhea. (He kept a mini-fridge with him in Thailand where he could chill his stool.)
There’s a similar story in his mouth–although it involves different species that are adapted to the conditions there.
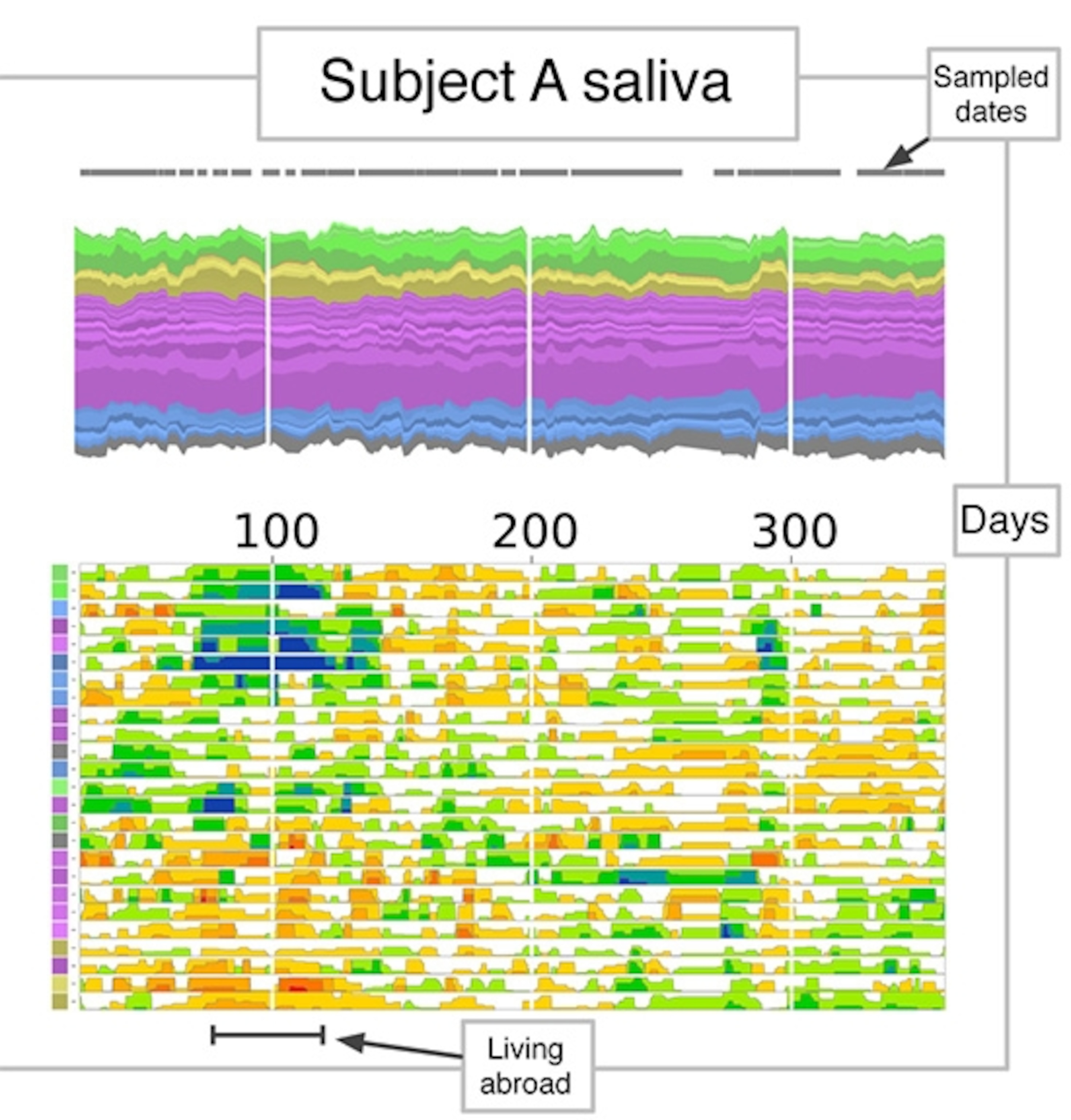
Now here’s Alm (he didn’t manage to go the whole year). The first thing you’ll notice is that Alm and David are microbially different. Over the year, they are consistently home to different kinds of bacteria.
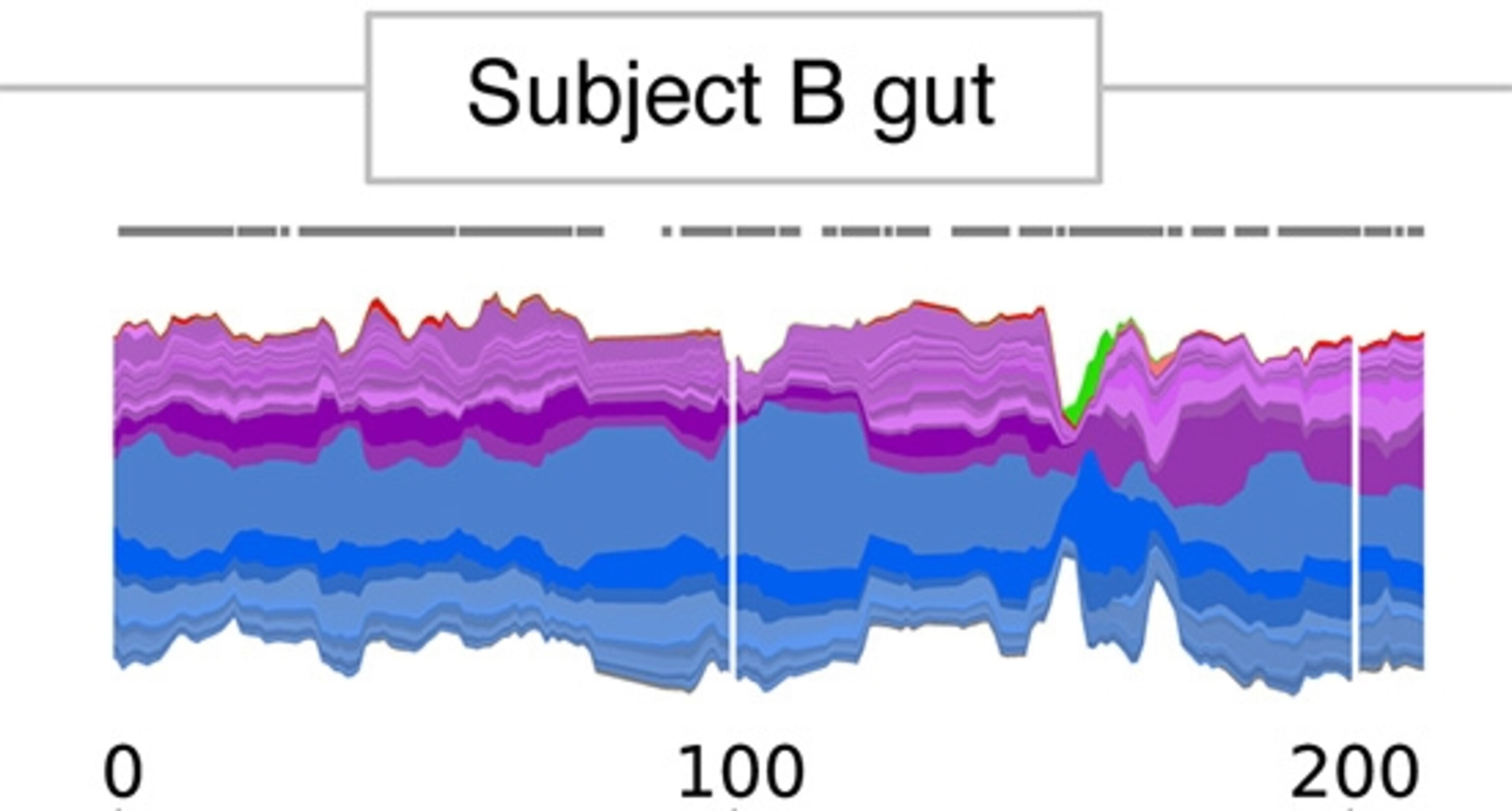
A startling pinch occurs around day 150. There’s a story behind that. On an unlucky visit to a restaurant, Alm got food poisoning–Salmonella to be specific. In fact, Alm was able to diagnose himself through his experiment even when his doctor was assuring him he had a virus. The infection was so intense that on one day, Salmonella made up 29.5% of all the DNA fragments he discovered in his stool. Fortunately, his food poisoning wasn’t bad enough to require antibiotics.
In the more detailed graph below, you can see that even after Salmonella vanished from his gut, his microbiome remained different.
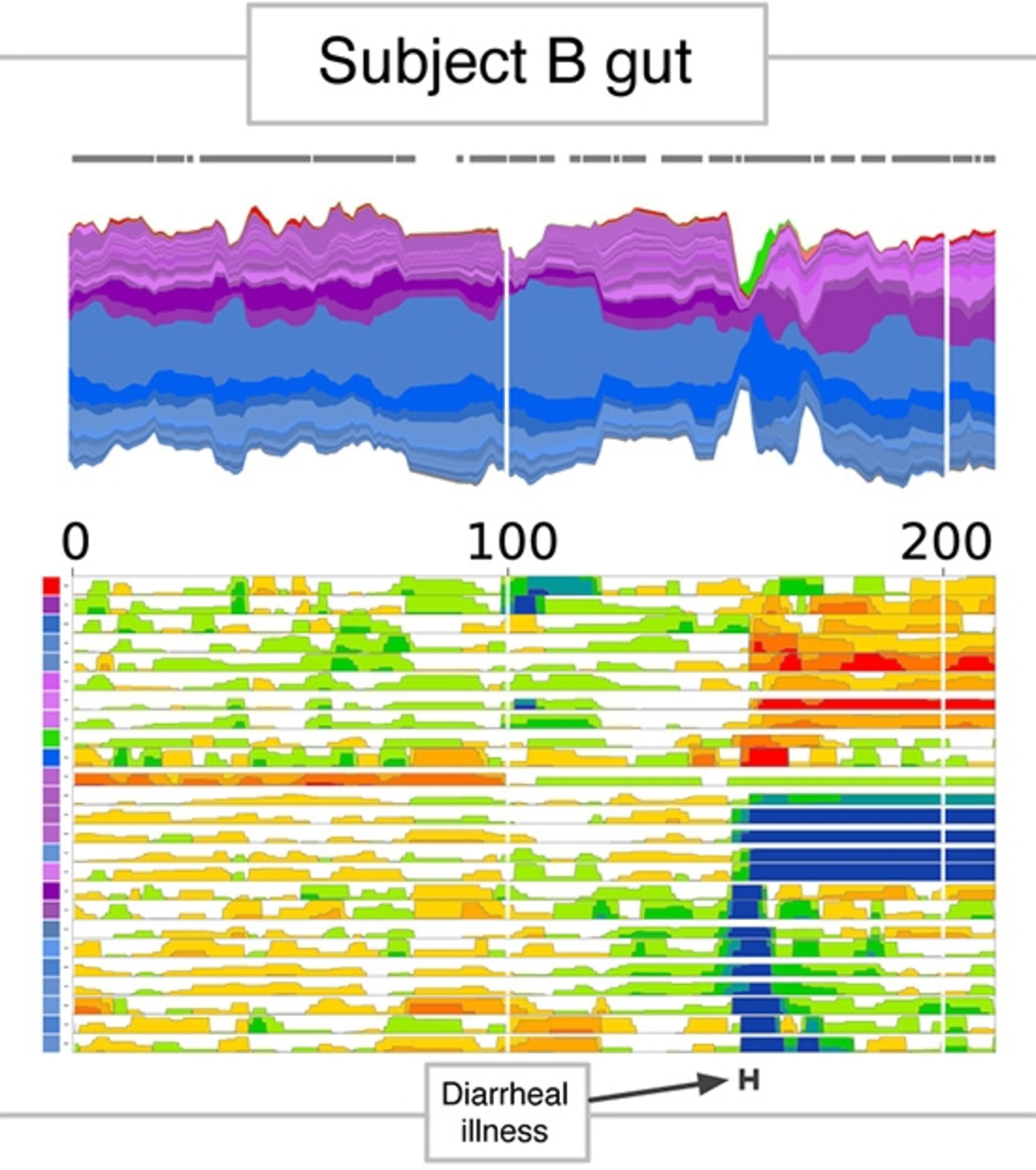
Looking at their microbiomes as just isolated species only took Alm and David so far, though. In reality, our microbes exist in communities. They may be adapted to the same acidity and other conditions, or they may depend on each other for survival.
Alm, David, and their colleagues found a way to visualize these communities as well. They identified five clusters of species whose changes tracked closely together. Here’s a chart of the changes they underwent, with David in the top graph and Alm in the lower one.
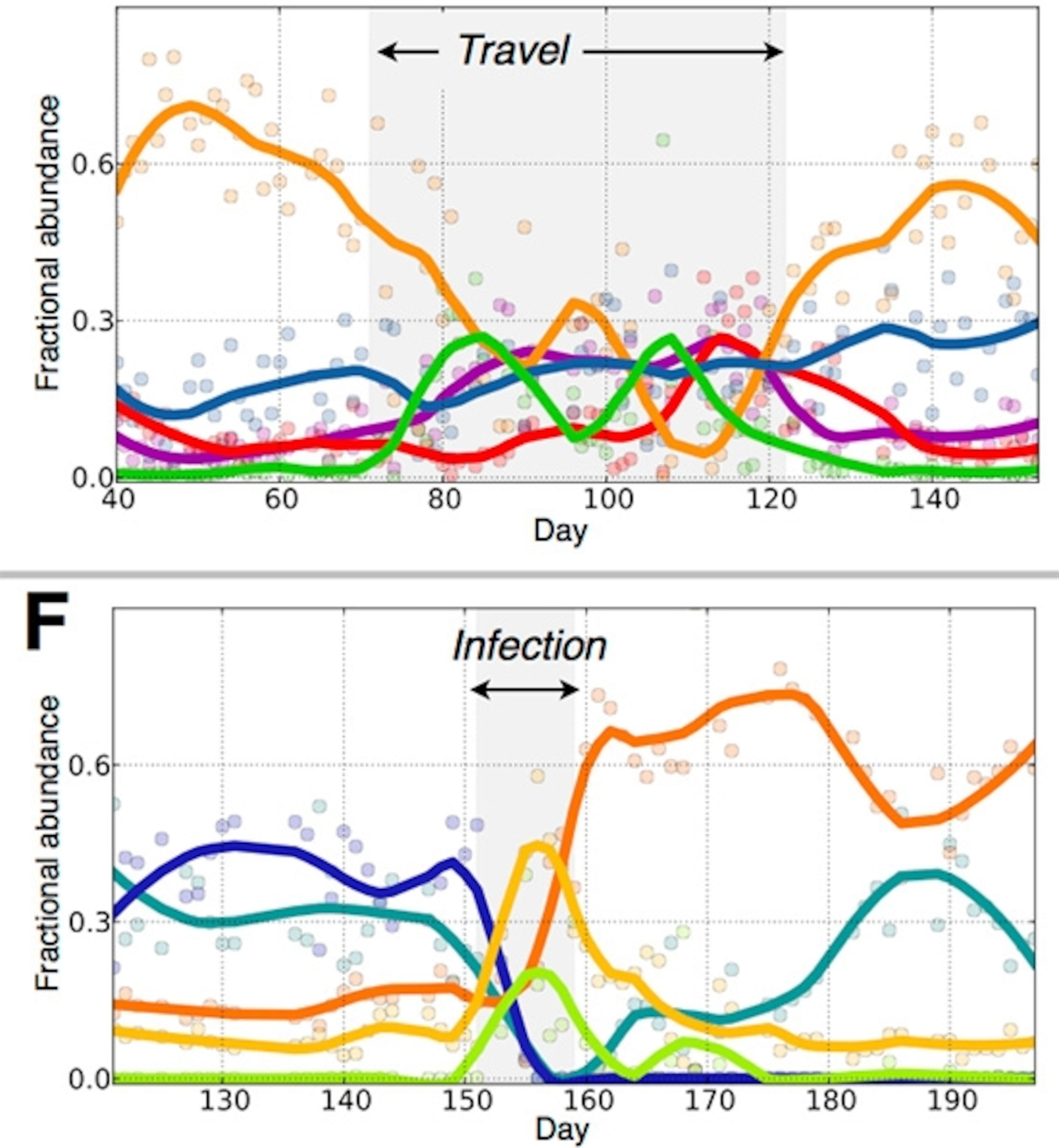
It appears that David’s microbiome switched from one state to another on his trip, and then back again. Alm’s microbes had a different experience. His food poisoning disrupted his microbiome, allowing a different combination of species to become dominant. It pushed him into a different state–a healthy one–and he’s now stuck there.
Alm, David, and their colleagues also tracked many other features of their lives during the study, from their meals to their moods. But aside from David’s trip to Bangkok and Alm’s bad dinner, they found little correlation between their experiences and their biodiversity. The only other factor that stood out was when David ate fiber-rich food–certain species of bacteria thrived the day after those meals.
It’s not possible to draw broad lessons from a two-person experiment. And it’s hard to imagine how it will become much bigger any time soon–who among us is ready to store stool away every day for a year? Perhaps someday someone will invent a lab-on-a-chip that you can swallow, and it swim in your gut, quietly monitoring your microbiome. At that point, I for one would happily track my inner zoo.